
Synthetic community [SynCom] transfer for the...
Tuberculosis, caused by bacteria in the Mycobacterium tuberculosis complex (MTBC), is a major global public health burden.
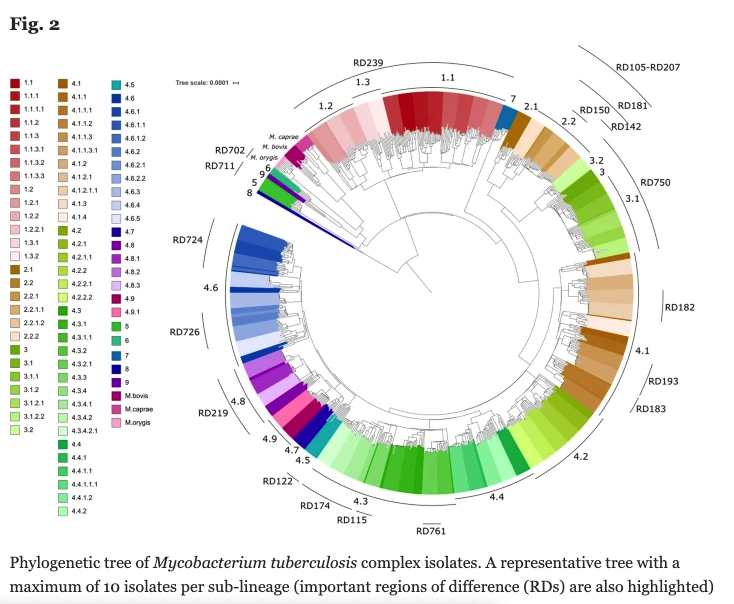
Strain-specific genomic diversity in the known lineages of MTBC is an important factor in pathogenesis that may affect virulence, transmissibility, host response and emergence of drug resistance. Fast and accurate tracking of MTBC strains is therefore crucial for infection control, and our previous work developed a 62-single nucleotide polymorphism (SNP) barcode to inform on the phylogenetic identity of 7 human lineages and 64 sub-lineages.